Haemophilia A is an X-linked disorder with an incidence of 1 in 5000 males. The common molecular defects in the F8 gene causing severe Haemophilia A are intron 22 and intron 1 inversions responsible for 45-50% of cases. Point mutations and gross deletion/insertions account for the remaining cases which are screened using DNA sequencing and fragment analysis. Genetic testing confirms the diagnosis and can aid in prenatal testing and counselling.
Intron 22 (int22) inversion
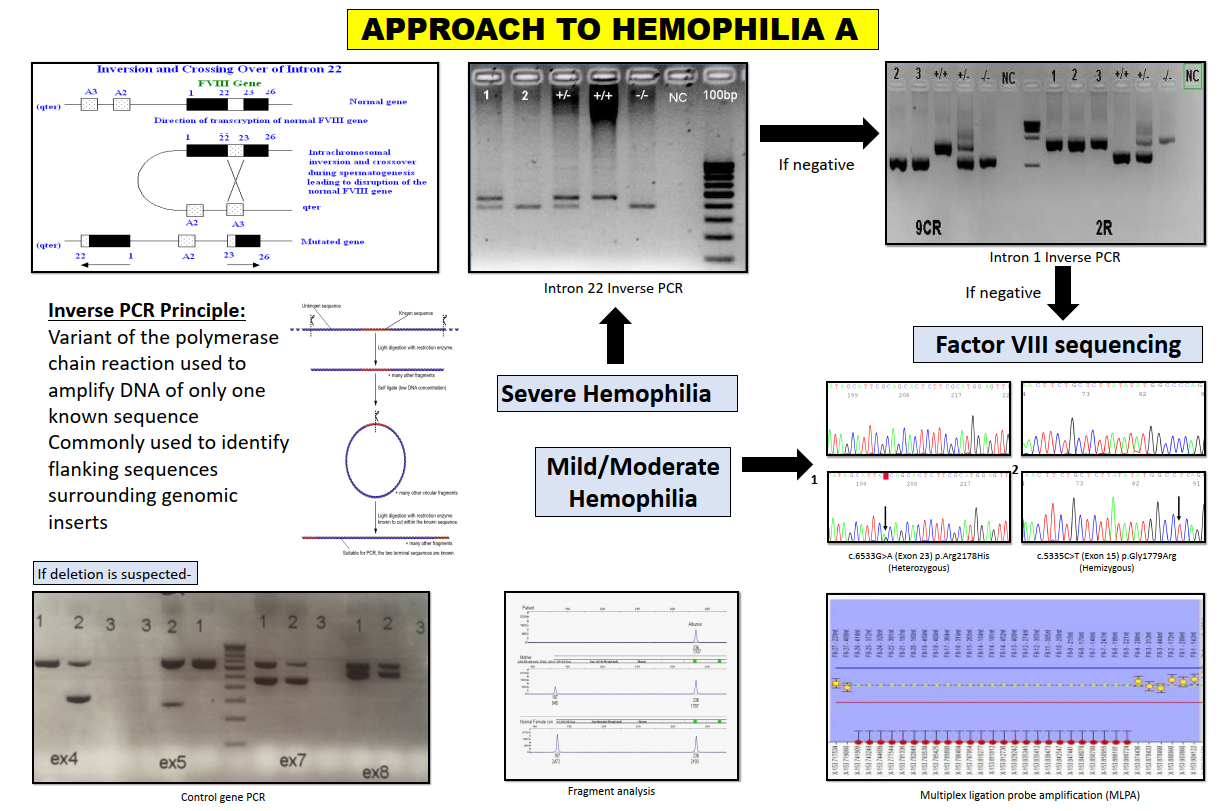
The F8 gene int22 inversion arises from homologous recombination between int22h-1, a region within intron 22 of F8, and one of two additional copies of the int22h-1 region. A technique to detect these inversions is Long distance PCR. In our lab, we perform inverse PCR for the detection of int22 inversion which involves three main steps (See reference 1):
- Restriction enzyme digestion of DNA
- Self-ligation of digested products to make DNA rings
- PCR amplification of DNA rings
Intron 1 (int1) inversion
The F8 gene int1 inversion arises from homologous recombination between int1h-1, a region within int1 of the F8 gene, and one additional copy of the int1h-2 region located approximately 140kb telomeric to the F8 gene. Different sets of primers are used to amplify int1h-1 and int1h-2 copies. Patients with the inversion will give different sizes of PCR products to those without the inversion (See reference 2).
If no inversions are detected, F8 gene sequencing is carried out to detect missense variants or small deletions/insertions/indels. If negative, large deletions and duplications are screened by gene dosage PCR, followed by fragment analysis and confirmed by multiplex ligation dependent probe amplification (MLPA).
